What is Gene Ontology?
Gene Ontology is a free online database that collects and combines information on gene function from a great number of species. [1] GO was born out of a need for consistent language describing genetic function, as well as a desire for an easy method of cross-referencing genetic data for homologous genes across species. [2] It is a vast resource for discovering homologs, genes with similar functions, and potential genetic interactions. Gene Ontology established three domains for describing the function of genes and gene products: biological process, cellular component, and molecular function. [1]
1. 'Biological Process' refers to pathways a gene/gene product is involved in. [1] This pathways are defined as having a defined beginning and end and being important to the function or continued survival of a biological unit. [2] 'Photosynthesis' would be an example of a biological process a gene product could be involved in.
2. 'Cellular Component' refers to the region (be it intra- or extracellular) in which a particular gene product is active. [1,2] For example, a DNA polymerase is active in the nucleus of a cell.
3. 'Molecular Function' refers to a gene product's specific biochemical function. [1] 'Ion channel' could be an example of a gene product's molecular function.
1. 'Biological Process' refers to pathways a gene/gene product is involved in. [1] This pathways are defined as having a defined beginning and end and being important to the function or continued survival of a biological unit. [2] 'Photosynthesis' would be an example of a biological process a gene product could be involved in.
2. 'Cellular Component' refers to the region (be it intra- or extracellular) in which a particular gene product is active. [1,2] For example, a DNA polymerase is active in the nucleus of a cell.
3. 'Molecular Function' refers to a gene product's specific biochemical function. [1] 'Ion channel' could be an example of a gene product's molecular function.
GLI3 at the Biological Level
|
GLI3 proteins are a component of the Hedgehog pathway, an early embryonic development mechanism. [3] They are post-transcriptionally modified into states that either repress or enhance the transcription of a number of proteins known to be involved in organogenesis. [3] The ratio of different post-transcriptionally-modified varieties of GLI3 is specifically linked to the formation of limbs and digits. [4]
|
GLI3 at the Cellular Level
GLI3 at the Molecular Level
|
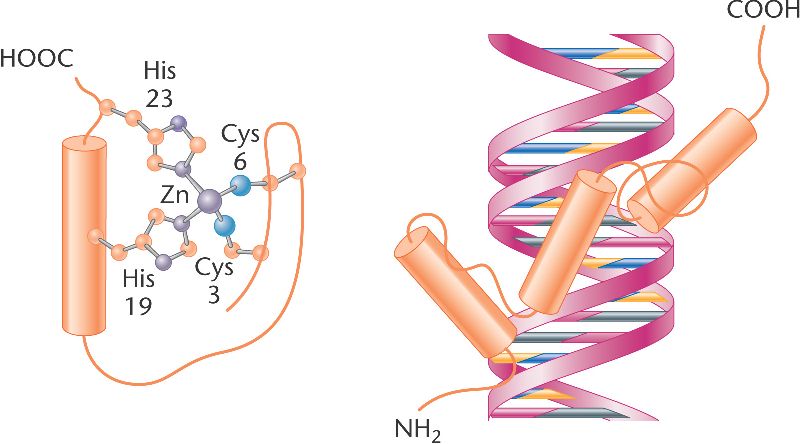
The GLI3 protein is a zinc-finger transcription factor. [4] It has a series of peptide sequences that bind to a segment of DNA, which enhances or represses the expression of a number of genes depending on where the transcription factor is attached. [6]
|
A zinc-finger domain binding to a segment of double-stranded DNA
|
Conclusion
A solid knowledge of GLI3 with regards to all three of Gene Ontology domains will be essential to determining its role in the mechanism determining digit number and placement. There is sparse research, currently, explaining the specific molecular function of GLI3, so discovering more about the protein on a molecular level should be a focus for further research into the role of GLI3.
References
- Gene Ontology. (n.d.). An Introduction to the Gene Ontology. <ftp://ftp.geneontology.org/go/www/GO.doc.shtml>
- Ashburner, M., Ball, C. A., Blake, J. A., Botstein, D., Butler, H., Cherry, J. M., Davis, A. P., Dolinski, K., Dwight, S. S., Eppig, J. T., Harris, M. A., Hill, D. P., Issel-Tarver, L., Kasarskis, A., Lewis, S., Matese, J. C., Richardson, J. E., Ringwald, M., Rubin, G. M., … Sherlock, G. (2000). Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature genetics, 25(1), 25-9.
- Carballo, G. B., Honorato, J. R., de Lopes, G. P. F., & Spohr, T. C. L. d. S. e. (2018). A highlight on Sonic hedgehog pathway. Cell Communication and Signaling, 16(1), 11. doi:10.1186/s12964-018-0220-7
- Büscher, D., Bosse, B., Heymer, J., & Rüther, U. (1997). Evidence for genetic control of Sonic hedgehog by Gli3 in mouse limb development. 62(2), 175-182.
- Cheng, F., Ke, X., Lv, M., Zhang, F., Li, C., Zhang, X., . . . Li, S. (2011). A novel frame-shift mutation of GLI3 causes non-syndromic and complex digital anomalies in a Chinese family. 412(11), 1012-1017.
- Fedotova, A. A., Bonchuk, A. N., Mogila, V. A., & Georgiev, P. G. (2017). C2H2 Zinc Finger Proteins: The Largest but Poorly Explored Family of Higher Eukaryotic Transcription Factors. Acta naturae, 9(2), 47-58.
Images
Header: https://phys.org/news/2017-09-internal-clock-human-cells.html
Image 1: http://geneontology.org/
Image 2: https://www.researchgate.net/figure/The-mammalian-Hh-signaling-pathway-Hedgehog-ligands-Sonic-hedgehog-Shh-Indian_fig6_235125014
Image 3: https://sites.nationalacademies.org/cs/groups/pgasite/documents/webpage/pga_049911.pdf
Image 4: http://bio3400.nicerweb.com/Locked/media/ch17/activator_zinc-finger.html
Image 1: http://geneontology.org/
Image 2: https://www.researchgate.net/figure/The-mammalian-Hh-signaling-pathway-Hedgehog-ligands-Sonic-hedgehog-Shh-Indian_fig6_235125014
Image 3: https://sites.nationalacademies.org/cs/groups/pgasite/documents/webpage/pga_049911.pdf
Image 4: http://bio3400.nicerweb.com/Locked/media/ch17/activator_zinc-finger.html
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.