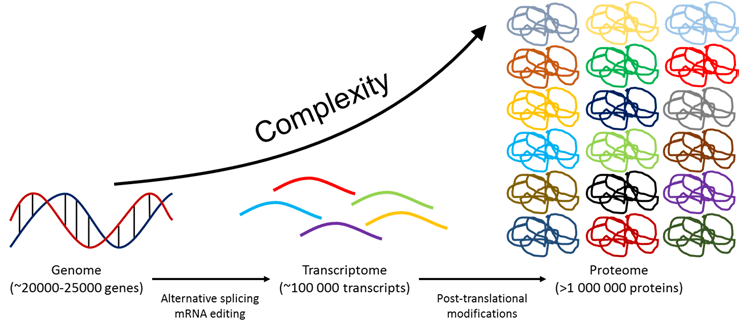
What are Post-Translational Modifications?
The term 'post-translational modification' refers to the extremely broad range of structural changes that proteins can undergo following transcription. [1] Using post-translational modifications, the same genetic sequence can code for a great number of functionally-distinct protein products. This explains how there are magnitudes more proteins in the human proteome than there are genes in the human genome. [2] Post-translational modifications vary across tissues, and can be permanent or reversible. [1] It is essential to consider the effects of post-translational modifications at all stages of a protein's existence when studying its activity, as they can change at any point. [1]
Post-translational modifications are detected via mass spectrometry, a chemical analysis technique which can be interpreted to discover chemical structures and differences in mass between samples.
Post-translational modifications are detected via mass spectrometry, a chemical analysis technique which can be interpreted to discover chemical structures and differences in mass between samples.
Because of alternative splicing and post-translational modification, the proteome is magnitudes larger than the genome from which it originates.
What is Protein Phosphorylation?
Phosphorylation is the act of adding negatively-charged phosphate groups to certain sites on a protein's sequence, which alter the protein's function. [2] Oftentimes, phosphorylation/dephosphorylation will act as an 'on/off' switch for a protein, activating and disabling it as needed. [2] The most common amino acid to be phosphorylated is serine, followed by threonine and then tyrosine. [2]
Are GLI3 Phosphorylation Sites Conserved Across Homologs?
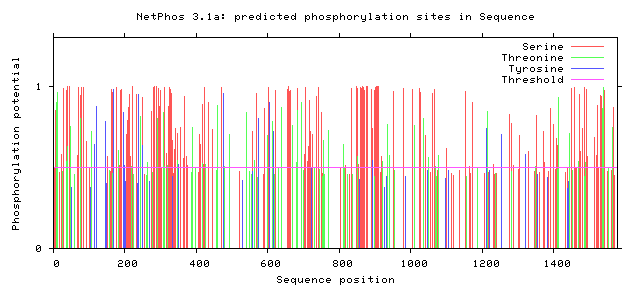
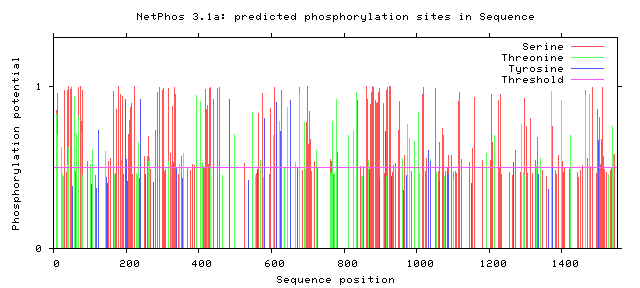
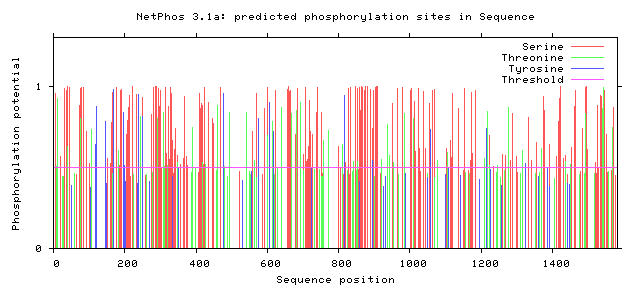
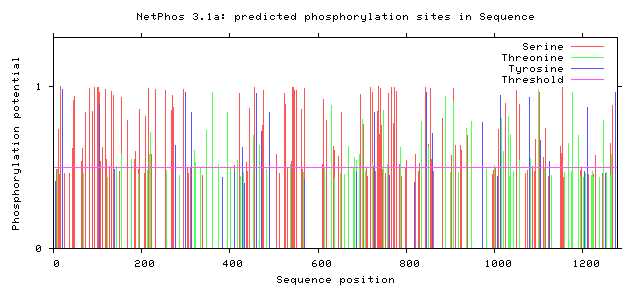
The following figures depict computationally-predicted phosphorylation sites on the sequence of GLI3 homologs for four species. The sites are predicted for the four most commonly-phosphorylated amino acids, and were calculated by NetPhos software.
Conclusions
It has been documented that GLI3 becomes activated by undergoing phosphorylation, although there does not appear to be a concrete consensus in the literature about where exactly this essential phosphorylation occurs. [3] From the comparison between the human GLI3 protein's phosphorylation patterns as well as those of its homologs, there do appear to be some immediately-apparent conserved regions. Most prominently, there appears to be a large and well-conserved 'island' of serine phosphorylation between 800 and 900 amino acids into the sequences. This is a promising lead, and further investigation is warranted. Understanding the activation patterns of GLI3 could be a fantastic clue towards understanding its role in determining digit number and placement.
References
- Duan, G., & Walther, D. (2015). The roles of post-translational modifications in the context of protein interaction networks. PLoS computational biology, 11(2), e1004049. doi:10.1371/journal.pcbi.1004049
- Overview of Post-Translational Modifications (n.d) Retrieved from: https://www.thermofisher.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
- Carballo, G. B., Honorato, J. R., de Lopes, G. P. F., & Spohr, T. C. L. d. S. e. (2018). A highlight on Sonic hedgehog pathway. Cell Communication and Signaling, 16(1), 11. doi:10.1186/s12964-018-0220-7
Images
Header: https://www.biocompare.com/Editorial-Articles/331118-Interrogating-Protein-Phosphorylation-and-PTMs-with-Mass-Spec/
Image 1: https://www.researchgate.net/figure/The-human-proteome-contains-many-more-species-compared-to-the-human-genome-making-it_fig4_313535513
Image 1: https://www.researchgate.net/figure/The-human-proteome-contains-many-more-species-compared-to-the-human-genome-making-it_fig4_313535513
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.