What is Transcriptomics?
Transcriptomics refers to the analysis of the 'transcriptome,' a compilation of all coding and non-coding RNAs that are in a cell at a given time and under a given set of circumstances. [1] The transcriptome can be studied using methods that focus on a single gene's expression (such as RT-qPCR and in situ hybridization) or via broad, high-throughput methods including microarray analysis and RNA sequencing, which can examine the expression of lots of genes at the same time. [1] Experiments dealing with transcriptomics are often used to examine how the expression of a particular gene or set of genes changes in response to development, maturity, and environmental factors. [1]
Common Transcriptomic Techniques
|
Microarray Analysis
This type of analysis is most often used to determine if a particular gene or set of genes is being expressed in cells and to what extend under particular conditions. [2] The technique is performed by generating complementary DNA (cDNA) from the RNA of a cell that is under specific conditions (developmental and environmental). [2] The cDNA is run across a chip that has had segments hundreds of genes of interested ligated to it in known locations and tagged with fluorescent proteins. [2] If any of the genes of interest were expressed in the cell at the time of cDNA generation, then the cDNA attaches itself to the gene segment on the chip and fluoresces. [2] |
|
RNA sequencing
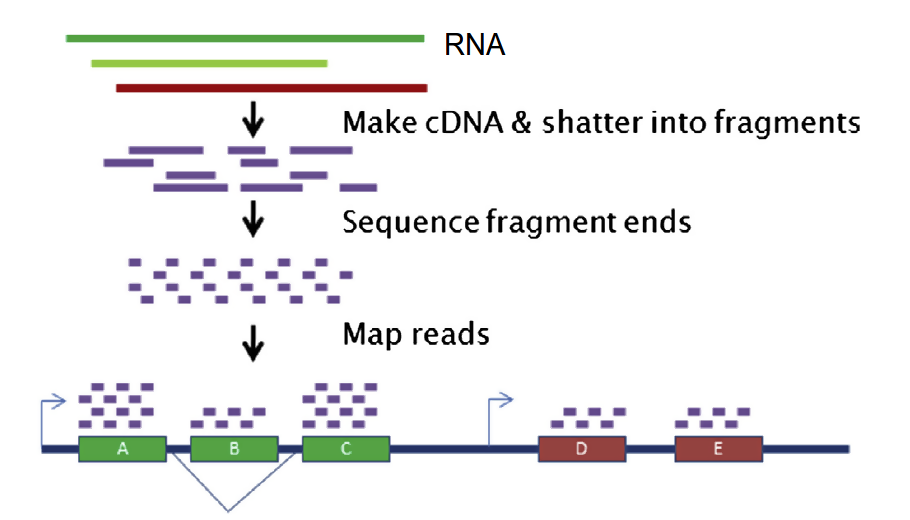
RNA-seq is a powerful modern technique that allows for whole-cell and whole-tissue transcriptomic analysis. [3] A cell's mRNA is used to generate cDNA, and fluorescent probes are attached to the cDNA fragments. [3] The fragments are sequenced and aligned against the genome. [3] This theoretically gives the expression levels for every portion of the genome under a particular set of circumstances. |
Transcriptomic Analysis of MYH7
|
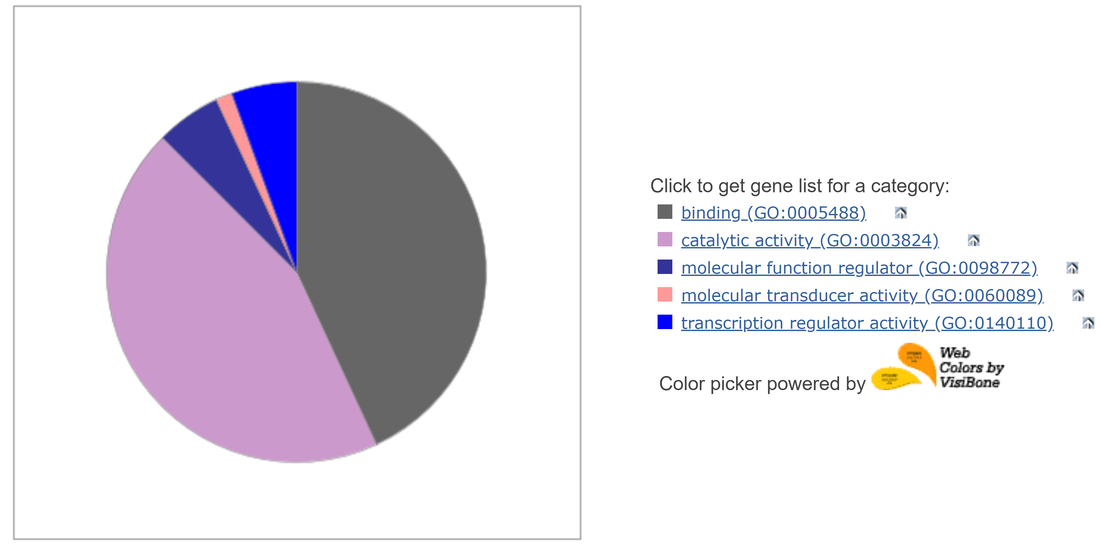
GLI3 is expressed in embryoid bodies, but not in embryonic stem cells
In a transcriptomic study conducted on mice embryonic stem cells and embryoid bodies, it was found that GLI3 is only expressed once an embryoid body is formed. The majority of genes clustered around GLI3 in the RNAseq analysis, when analyzed using Panther, were found to be involved in binding and catalytic activity. |
|
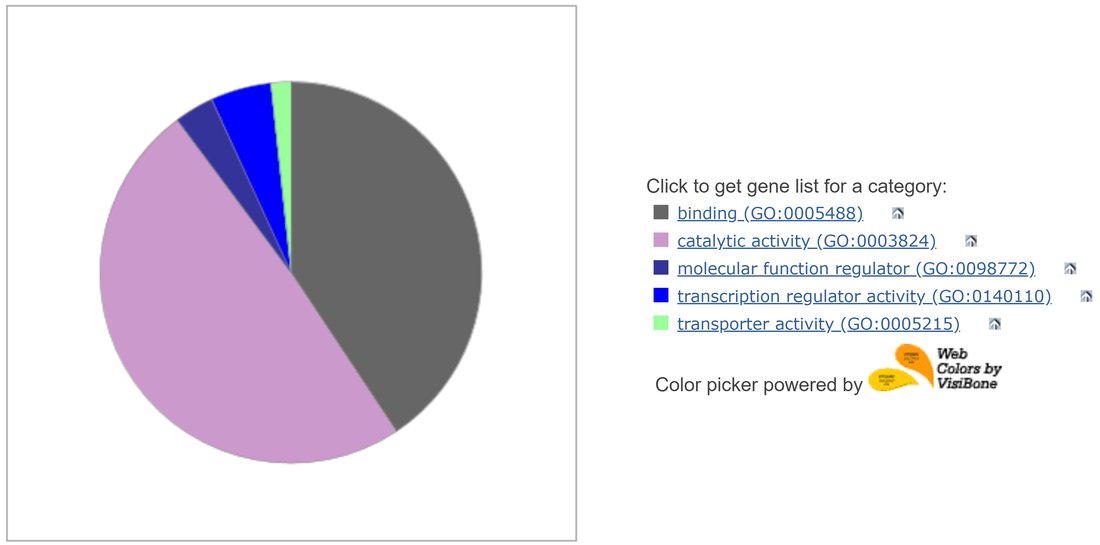
Downregulation of Hedgehog diminishes GLI3 expression
A transcriptomics study conducted on mouse embryoid bodies exposed to retinoic acid, an agonist of the Hedgehog gene, found that GLI3 expression was downregulated alongside Hedgehog. A Panther analysis of the genes clustered around GLI3 revealed that a majority of them were involved in binding and catalytic activity. |
Conclusion
GLI3 is known to be an essential transcription factor in embryonic development, which implies that there is a strong temporal factor in its expression. [4] These findings support this, as GLI3 is only upregulated once an embryoid body is formed and embryonic development begins. The genes most closely clustered around GLI3 are consistently found to be involved in binding and catalytic activity, which could imply a regulatory role in DNA expression similar to GLI3 itself. The complex web of transcription factors involved in early development are a promising lead in determining the mechanism behind digit development, and call for more specific experiments in that field, as literature is sparse concerning expression levels in limb buds during early development.
Interestingly, GLI3 is reported to be downregulated as a result of Hedgehog expression being hindered by retinoic acid. This information appears to disagree with reports that GLI3 and Hedgehog have a relationship of antagonistic expression. [5]
Interestingly, GLI3 is reported to be downregulated as a result of Hedgehog expression being hindered by retinoic acid. This information appears to disagree with reports that GLI3 and Hedgehog have a relationship of antagonistic expression. [5]
References
- Lowe, R., Shirley, N., Bleackley, M., Dolan, S., & Shafee, T. (2017). Transcriptomics technologies. PLOS Computational Biology, 13(5), doi:10.1371/journal.pcbi.1005457
- Microarray. (n.d.) Retrieved on 3/4/2019 from https://www.nature.com/scitable/definition/microarray-202
- Wang, Z., Gerstein, M., & Snyder, M. (2009). RNA-Seq: a revolutionary tool for transcriptomics. Nature reviews. Genetics, 10(1), 57-63.
- Patel, R., Singh, C. B., Bhattacharya, V., Singh, S. K., & Ali, A. (2016). GLI3 mutations in syndromic and non-syndromic polydactyly in two Indian families. Congenital Anomalies, 56(2), 94-97. doi:10.1111/cga.12139
- Büscher, D., Bosse, B., Heymer, J., & Rüther, U. (1997). Evidence for genetic control of Sonic hedgehog by Gli3 in mouse limb development. 62(2), 175-182.
Images
Header: https://www.the-scientist.com/news-opinion/how-statistics-weakened-mrnas-predictive-power-31484
Image 1: https://rep.bioscientifica.com/view/journals/rep/130/1/1300001.xml
Image 2: https://www.healthcarebusinesstoday.com/here-are-all-the-steps-involved-in-the-rna-seq-data-analysis/
Image 1: https://rep.bioscientifica.com/view/journals/rep/130/1/1300001.xml
Image 2: https://www.healthcarebusinesstoday.com/here-are-all-the-steps-involved-in-the-rna-seq-data-analysis/