What are Protein Interaction Networks?
Protein interaction networks are visual representations of the web of protein-protein interactions within cells and tissues of an organism. [1, 2] These interactions are specific and of biological importance, but the exact manner of interaction can vary. Examples of such interactions include being a component of a signaling cascade or a post-translational modification of one or both proteins. [1, 2] These interactions can be dynamic (lasting only briefly) or constant (a long-term or permanent binding/interaction). [1, 2]
Protein-protein interactions can be determined experimentally, or predicted using computing software. This compiled information can then be used to create a protein interaction network. The network can then be analyzed to find patterns in the functions or localization of the interacting proteins.
Protein-protein interactions can be determined experimentally, or predicted using computing software. This compiled information can then be used to create a protein interaction network. The network can then be analyzed to find patterns in the functions or localization of the interacting proteins.
GLI3 Interaction Network
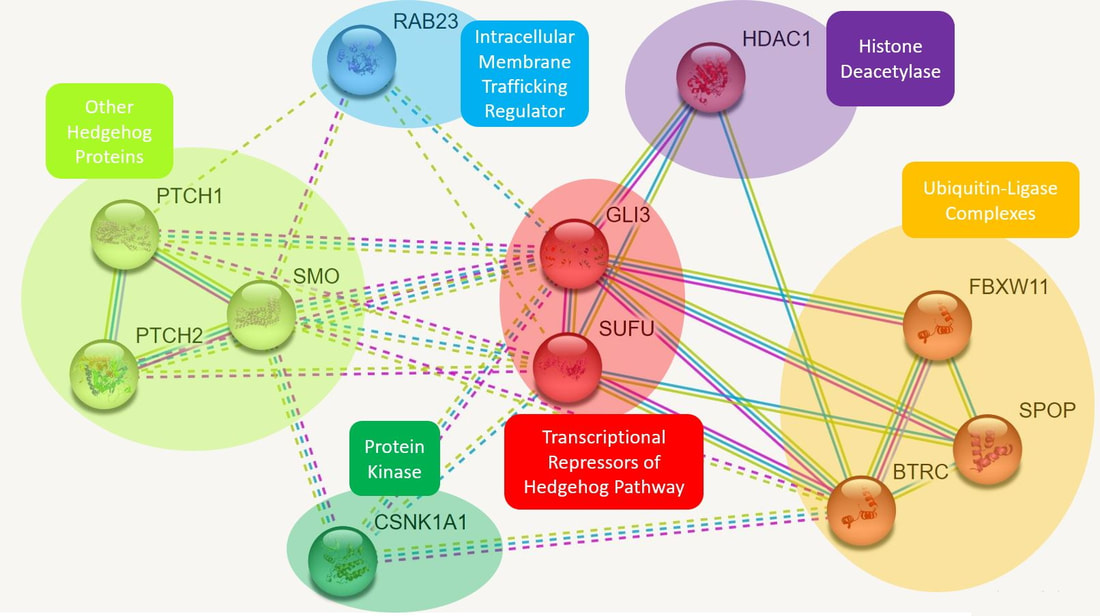
Figure 1. A protein network for GLI3 generated using the STRING database. Proteins have been sorted into categories roughly defined by their functions.
Conclusion
There are a few oddities to note from the STRING-generated protein network for GLI3. Firstly, while GLI3 is displayed as interacting with a number of proteins in the Hedgehog pathway, it is not shown directly interacting with the Hedgehog gene, despite the literature detailing the GLI3-Hedgehog gradient and their antagonistic relationship. This suggests that there is either an unknown interaction between the proteins (perhaps localized solely to limbs), or there are intermediary proteins involved in the pathway.
The next protein of interest is RAB23, which is known to be a regulator of intracellular trafficking. This is likely the protein controlling GLI3's movement into the nucleus, and seems to be a promising lead to learning more about GLI3's expression and activity as a repressor.
Finally, GLI3 is shown to interact with three proteins in ubiquitin-ligase complexes, which ubiquitinate proteins to mark them for degradation via proteasome. This seems to be an active component of GLI3's regulation, and is worth investigating further.
The next protein of interest is RAB23, which is known to be a regulator of intracellular trafficking. This is likely the protein controlling GLI3's movement into the nucleus, and seems to be a promising lead to learning more about GLI3's expression and activity as a repressor.
Finally, GLI3 is shown to interact with three proteins in ubiquitin-ligase complexes, which ubiquitinate proteins to mark them for degradation via proteasome. This seems to be an active component of GLI3's regulation, and is worth investigating further.
References
- Protein-protein interaction networks. (n.d.) EMBL-EBI. Retrieved from: https://www.ebi.ac.uk/training/online/course/network-analysis-protein-interaction-data-introduction/protein-protein-interaction-networks
- Snider, J., Kotlyar, Max., et al. (2015). Fundamental of Protein Interaction Network Mapping. Retrieved from: http://msb.embopress.org/content/11/12/848#abstract-2
Images
Header: https://datascience.wisc.edu/academics/
This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.